Testing a Spatial Memory Index for Strap
One of the pieces I still need to build for strap is a long-term memory store. As I’ve written about before, strap compresses context to work within small token budgets. But context compression only handles what’s in the current conversation. The bigger question is how to surface relevant memories from past sessions.
The obvious approach is brute-force cosine similarity over sentence embeddings. It works for a small number of stored memories, and it’s fast enough. But I got curious whether a spatial index could do better, and whether you could make that index inspectable at the same time.
I spent a couple days running experiments with Claude Code and “we” wrote the results up as a paper. If you’ve been out of school for a while, this should be a pretty enlightening example as to why professors are having trouble telling actual student work from AI assisted papers. To be clear, I would never hand something like this in as my own work. In fact this is the first time I have tried anything like this, and I have to say, it’s impressive what it can produce with just some basic guidance.
The following is the short version of the paper and the experiments I’ve been doing the past few days, or you can check out the whole paper if you are into that kind of thing.
The Idea
Sentence embeddings from all-MiniLM-L6-v2 are 384-dimensional vectors. You can compress those to 3D using PCA or UMAP and build an octree over the projected coordinates. At query time: embed the query, project to 3D, navigate the tree to the right leaf, pull candidates, and then cosine re-rank on the full 384-dim vectors.
The reason I wanted an octree specifically is incremental insertion. You can add a new memory by walking root-to-leaf. Methods like HNSW need to wire the new node into a proximity graph across multiple layers. IVF assigns points to k-means centroids, and if the data distribution shifts, those assignments will degrade. For a memory store that’s updating continuously as an agent runs, I was thinking that property matters quite a bit.
The 3D constraint also means you can render the whole index as a scatter plot and look at it. I liked that idea.
Before any of that, I needed to confirm the embedding pipeline could run in a Go binary without an external service. The ONNX runtime works in golang, the INT8 quantized model is only 23MB, and the whole thing packs into a ~64MB binary with the runtime, model, and tokenizer included. Embedding quality was identical to float32 at four decimal places.
Small Scale: Promising
📝 Side note: Recall@5 means what fraction of a query’s 5 true nearest neighbours (computed by brute-force cosine on the full vectors) the pipeline returned. 100% means all five results matched.
On a 25-sentence pilot corpus the results looked good. PCA 16D plus a KD-tree got 100% recall@5 at about 35 microseconds per query transform. UMAP 3D got 94.4% recall, but at nearly a second per query, it kind of ruled it out for the harness.
PCA 3D alone got 90.4% which I was happy with. The variance numbers looked somewhat promising at: 27% of the embedding space explained by three components on a small, topically tidy corpus.
Large Scale: Not Promising
I scaled up the tests to 120,000 sentences from the AG News headlines data set to see how it held up.
It did not hold up.
The problem is PCA variance collapse. On 25 sentences, 3 components explain about 27% of the variance. On 120k diverse sentences, that drops to 8.9%. The 3D position of a point becomes almost uncorrelated with its semantic content. Routing a query to the right spatial cell is close to random. PCA at every dimensionality I tested (3D, 8D, 16D) came in under 1% recall.
UMAP is better because it explicitly preserves local neighbourhood structure instead of maximising global variance. At 120k sentences it got 32.8% recall at 24ms per query. t-SNE got 19.2% at 128ms.
Brute-force cosine got 100% recall at 1.15ms.
So the index is slower and less accurate. At this corpus size, scanning everything wins.
What the Octree Is Actually Good For
The 3D index turned out to be genuinely useful as a debugging tool during development. When PCA was the projection, I could look at the scatter plot and immediately see that semantically unrelated sentences were landing in the same spatial cells. That told me the routing was broken before I measured recall.
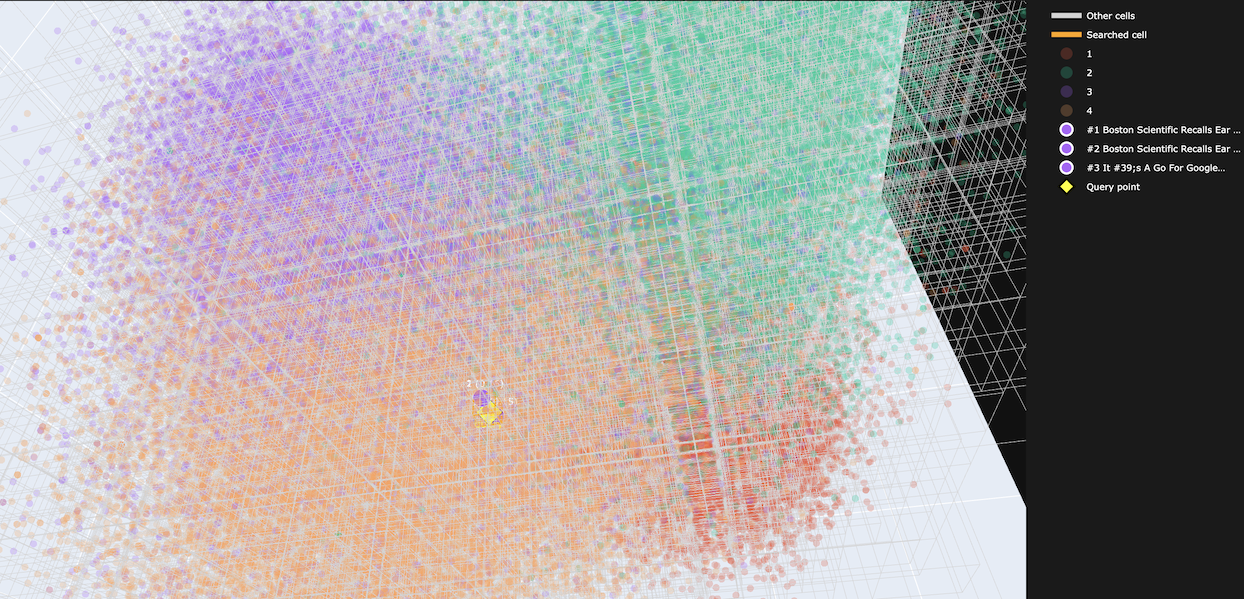
HNSW and IVF don’t give you that. Their internal structures are not something you can render and navigate interactively. The octree at 3D is inspectable by construction.
Where This Leaves Strap
For the actual memory store, brute-force cosine is probably the right answer until the corpus is large enough to be slow. At a few thousand memories it should still be sub-millisecond. The octree might still be worth keeping as a parallel visualisation structure so you can inspect what the model considers semantically similar at any point. But really, you can just generate that when ever you want.
The Go embedding pipeline works, and I’ll probably be using it in Strap unless I can find something better. The spatial indexing experiment is interesting, but it seems like I should just scan the vectors or use the well worn path of KD-Trees or maybe SphereTrees.
Full details in the paper and source code for everything can be found on github.